Folding@home lets us generate a lot more simulation data than is typically possible otherwise. Historically, lots of simulation studies have relied on a few simulations, let’s say one to ten. With Folding@home we can run thousands of trajectories. Each is reasonably long by the standards of the field but is typically short compared to the timescale of the dynamics we intend to study. By taking lots of shots on goal, these simulations give us statistics on slower dynamics that are typically beyond the reach of any one simulation.
Markov state models (MSMs) are the methodology that we and others have developed to make sense of the large number of trajectories that the Folding@home community generates.
MSMs are analogous to the way that your favorite maps app creator infers potential routes and speeds from the GPS coordinates of their users. In that case, they use GPS coordinates stored at regular intervals in time to infer where intersections are and the speed with which one can get between them. Most of us don’t drive very far. We’re typically just running to the store or commuting to work, not making cross country road trips. However, as long as there’s overlap between the intersections that we go through, the maps app creators can infer a map of the entire country (and beyond).
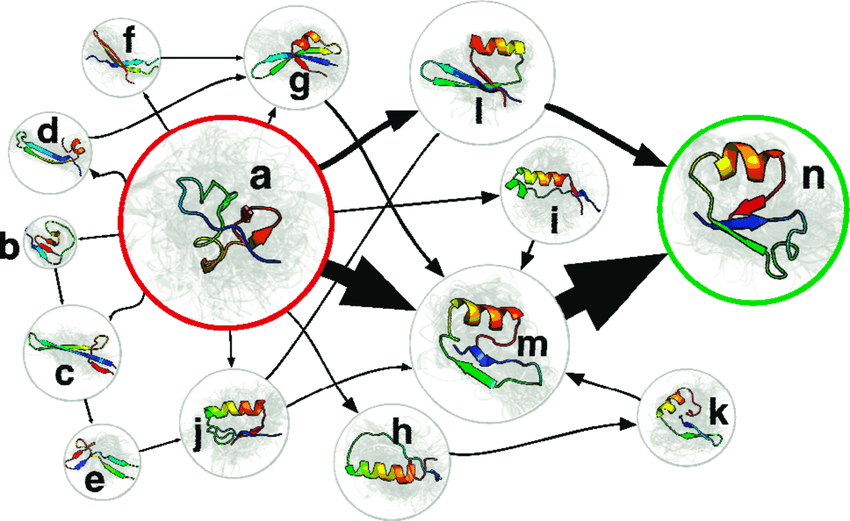
Likewise, MSMs build maps from a large number of simulations. Each simulation consists of protein structures stored at regular intervals in time, akin to GPS coordinates from a driver commuting to work. By taking advantage of protein conformations that are visited by multiple simulations, we’re able to identify common structures and the speeds of getting between them, akin to intersections and speeds of getting between them in our map analogy. Like a map, the MSM then provides a basis for asking all kinds of questions about what’s out there and how to navigate that space.
Like maps apps, MSMs are a mature technology with seemingly endless possibilities for improvement. We recently organized a special issue that brings together papers on recent advances in the construction and use of MSMs, which you can find here.