When it comes to designing novel drugs, achieving specificity is a major challenge. An effective drug must bind tightly to its target protein while avoiding unwanted side effects that can result from interactions with other proteins. This challenge becomes even more complex when targeting specific members of protein families with similar structures. Additionally, some enzymes share substrates, like ATP, across various protein families, making it difficult to design compounds that compete with endogenous ligands without causing off-target effects.
One innovative approach to drug design is targeting allosteric sites rather than active sites. Allosteric compounds can enhance desirable protein functions, offering a unique way to achieve specificity. These sites are often less conserved than active sites, making it easier to develop specific drugs. In recent years, highly specific allosteric compounds have been serendipitously discovered through high-throughput screens, targeting various proteins such as G-protein-coupled receptors, myosins, kinases, and β-lactamases. Despite these successes, designing drugs that target allosteric sites from scratch is challenging because experimental structural studies often provide limited insights into a protein’s conformational landscape.
One specific area of interest is myosins, a superfamily of ATPases that play crucial roles in various cellular processes. Myosins have the potential to be valuable drug targets for numerous diseases, but their complexity and the existence of multiple isoforms make targeting specific myosin variants extremely difficult. For instance, there are 38 myosin genes in the human genome, and individual cells express about 20 different myosin isoforms. Compounds like mavacamten have shown promise in clinical trials for heart-related conditions, but there is a need for more myosin modulators to address a broader range of diseases. However, the challenge lies in targeting specific myosin isoforms due to their highly conserved motor domain fold and active site structure.
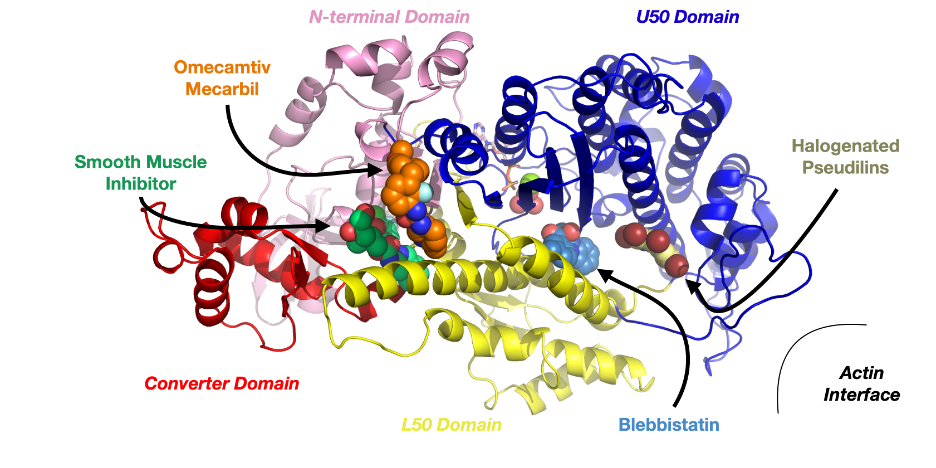
Figure caption: Structure of a myosin protein highlighting the binding sites of some known allosteric modulators, including blebbistatin.
Blebbistatin, a myosin-II specific allosteric inhibitor, has been a subject of study to understand the molecular mechanisms governing drug specificity. It was discovered in a high-throughput screen targeting nonmuscle myosin IIs and was found to broadly inhibit various myosin-II isoforms while sparing other myosin families. The key to its selectivity lies in the dynamics of the blebbistatin pocket and the conformations myosin isoforms adopt in solution.
Through all-atom molecular dynamics simulations, this study has shown that the probability of the blebbistatin pocket opening is higher in more sensitive myosin isoforms, which explains differences in drug potency. This finding, along with differences in the pocket’s residue composition, provides insights into the factors contributing to drug specificity. These results demonstrate the role of pocket dynamics and conformational selection in achieving drug specificity and highlight the potential for precision medicine through computational modeling.
In conclusion, the study of blebbistatin sheds light on the intricate world of drug specificity in the realm of myosin inhibitors. It emphasizes the importance of understanding the dynamic interplay between drug molecules and protein structures. This knowledge has the potential to open doors to more precise drug design, allowing us to target specific isoforms and improve the effectiveness of therapeutic interventions. As the field of precision medicine advances, computational modeling and simulations like the ones used in this study offer promising opportunities to tailor treatments to individual patients and address a wide range of diseases with unprecedented specificity.
