As the COVID Moonshot continues to make rapid progress, the fourth COVID Moonshot free energy calculation sprint focused on a backup lead series that suddenly pulls into the lead!

It’s been an incredibly busy time at Folding@home HQ, with a huge number of major projects all coming together at the same time: Between the CUDA-enabled OpenMM core (core22), a major update to the SARS-CoV-2 cryptic pockets paper, organization of major data releases, setting up some critical new projects, and having two of our amazing scientists (Hannah Bruce Macdonald and Matt Wittmann) making previously-planned moves to new positions, it’s been a hectic few weeks since our last update. But rest assured, the science has not stopped, and we’re making extremely rapid progress toward a new small molecule therapy for COVID-19.
As a reminder, you can keep up with the latest updates on the COVID Moonshot by subscribing to the email newsletter for frequent updates:
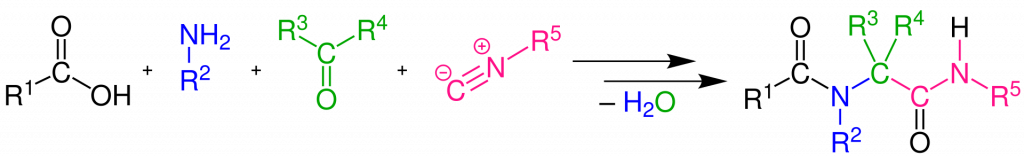
Sprint 4, which just wrapped up, focused on one of our back-up lead series, the Ugi series, an extremely interesting series championed by the London lab at the Weizmann Institute. Ugi reactions are particularly interesting for designing protease inhibitors because they can mimic the peptide backbone of the natural substrate peptide that the protease binds to tightly and cleaves, but still allow a wide variety of compounds to be synthesized using the same reaction, which brings together four different starting materials (which install interesting chemical groups at five distinct sites), creating a huge variety of possible compounds that can be synthesized. Even more surprisingly is that this can all happen in one step, in a single reaction vessel, allowing even robots to make these molecules very quickly and inexpensively!
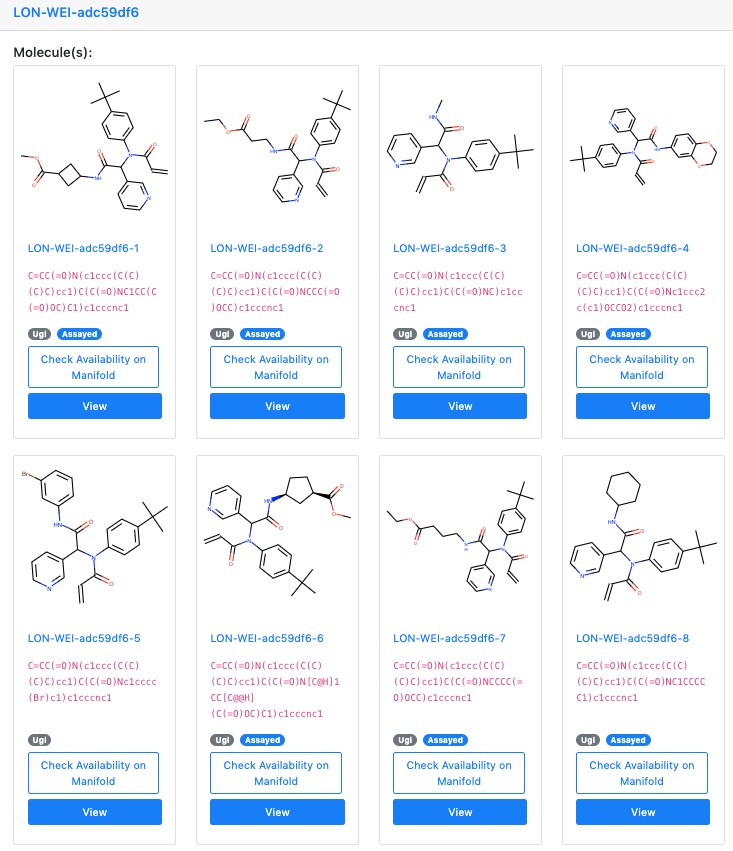
Some of the original compounds screened by the London Lab were a large library of compounds made with these Ugi reactions. The original goal was to develop a covalent inhibitor, so the library contained a large number of compounds with covalent warheads that could potentially form tight irreversible bonds with a particular cysteine residue (Cys145) in the SARS-CoV-2 main viral protease:
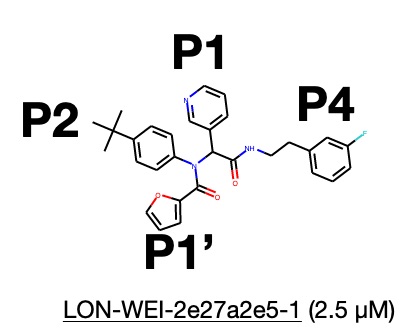
One of these compounds, LON-WEI-adc59df6-47, bound relatively well, with an IC50 of ~3.0 µM. The London lab then made several noncovalent version of this, LON-WEI-2e27a2e5, which—much to everyone’s surprise, worked even better, clocking in at an IC50 2.5 µM in the fluorescence assay (where lower is better since it means you need less drug to shut down 50% of the protease in the cell)! Since we already had three other major lead series (the 3-aminopyridines explored in Sprint 1 and 2, the benzotriazoles in Sprint 3, and the quinolones yet to feature in a sprint), we relegated the Ugis to a backup series that a few weeks of free energy calculations on Folding@home might help continue to optimize while we pushed full speed ahead on the three other main lead series.
Sprint 4: Optimization of Ugi P2 substituents
Starting from LON-WEI-2e27a2e5-1, our task in Sprint 4 was to identify which reactant we could swap out to optimize the substituent that fits into the P2 pocket.
Thanks to Matt Wittmann and Hannah Bruce Macdonald (and her analysis toolkit), we rolled out an updated science dashboard for Sprint 4, featuring a breakdown by compound, by microstate (since each compound can exist in a number of different protonation states or stereoisomers depending on the assay conditions and how they were synthesized), the individual free energy transformations we compute. We are also now integrating experimental data into the overall estimated affinities for each compound.
WCMC Pharmacology graduate student Melissa Boby sorted through the top compounds to build a final list of starting materials to buy to form the top-scoring compounds:
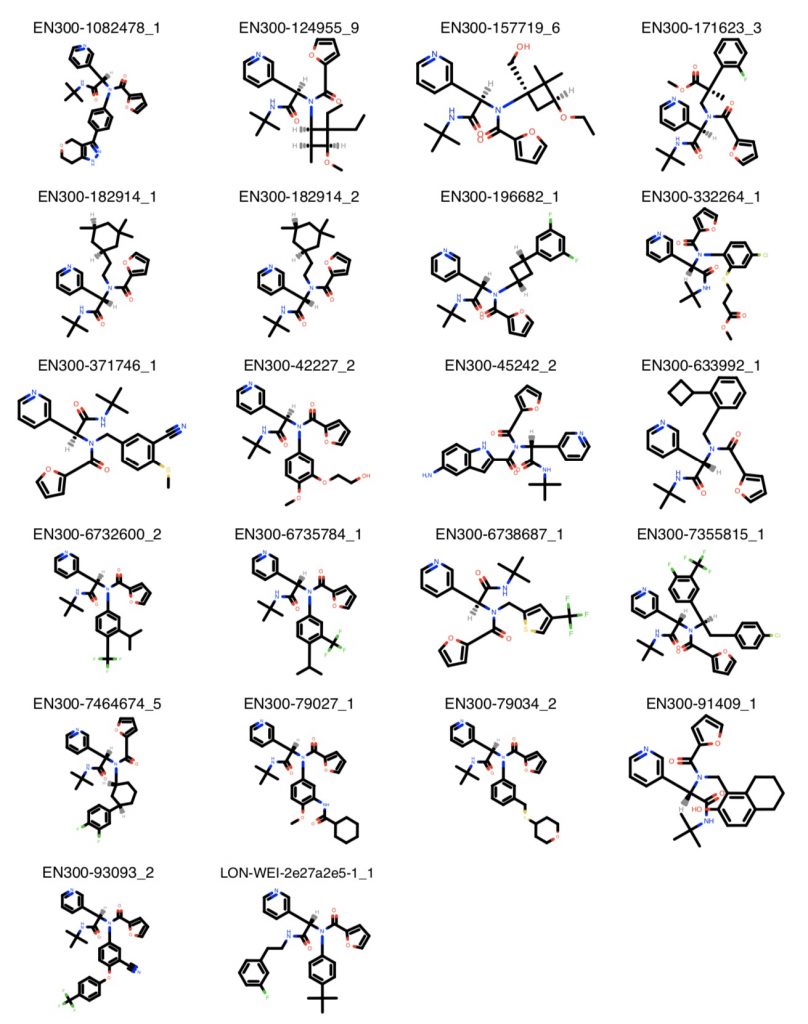
Just as we were doing this, something extremely unexpected happened: The chirally-separated version of the compound we started with turned out be 100 nM! That’s 25 times more potent (meaning you need 25x less drug to get the same effect in shutting down the protease) than what we thought we started from!
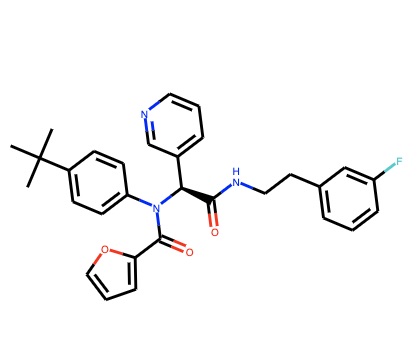
The exciting aspect of this is that we have apparently been working on an extremely potent compound series all along in Sprint 4, thinking it was “just” a backup series. All of the work we did has suddenly become much more important.
The starting materials we ordered from Sprint 4 have already arrived at the Weizmann, and the London lab is already hard at work synthesizing new compounds based on Sprint 4 designs. We’re excited to be able to share the results with you once they start trickling in.
More blogs to come in the next few days
Stay tuned, and keep folding! In the next few days, we’ll be blogging about
- Results from Sprints 1 and 2, where data has started to come in!
- How the process of small molecule drug discovery works, where the COVID Moonshot is in this process, and the steps left to bring the first potential drug to the clinic
- New exciting projects aimed at staving off drug resistance for COVID therapeutic antibodies
- And much more!
