The open science COVID Moonshot has made steady progress toward developing a new patent-free therapy for COVID-19. Folding@home has been working closely with the Moonshot (see our earlier post) during its initial phase of identifying potent compounds that inhibit the main viral protease (Mpro) of SARS-CoV-2:
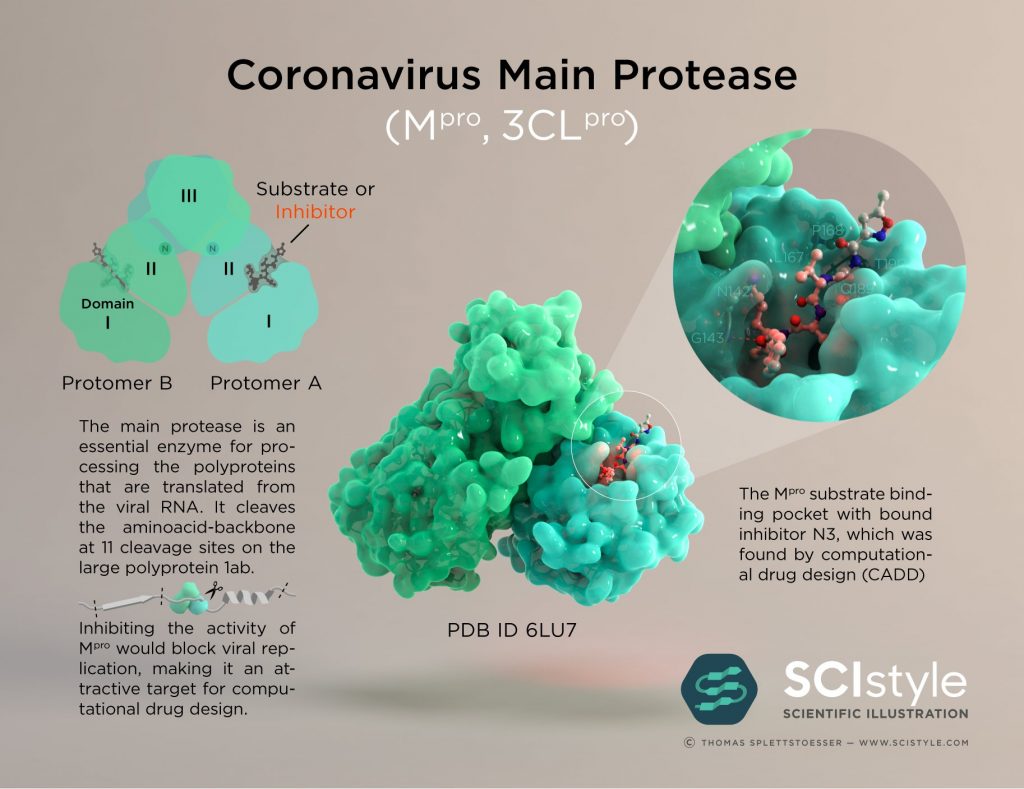
Up to now, the goal has been to find compounds that could be further refined into molecules with potent antivirals with good safety profiles. The progress has been incredibly rapid—over 800 compounds have been made and tested to date:
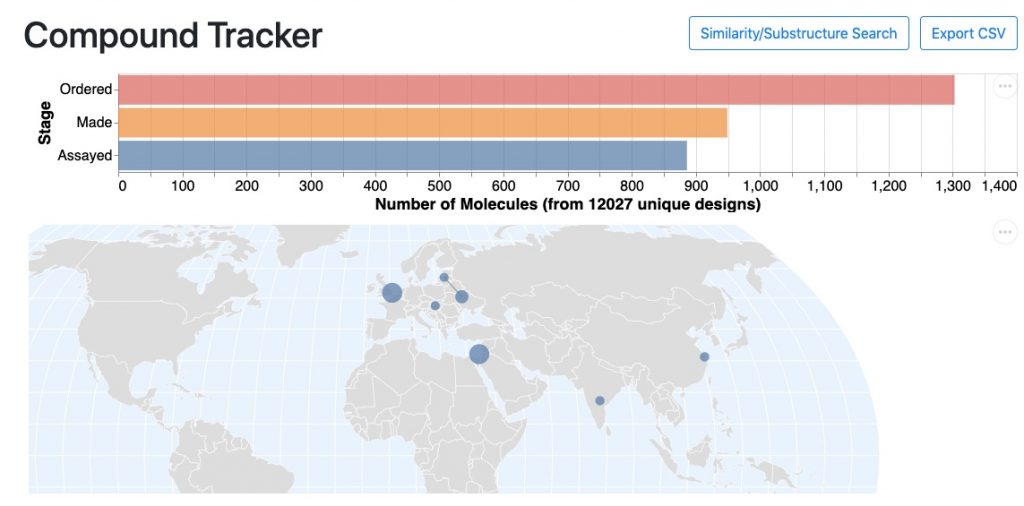
All the hard work has paid off—the Moonshot now has multiple lead compounds with biochemical potencies better than 1 micromolar, which makes them a great starting point for lead optimization.
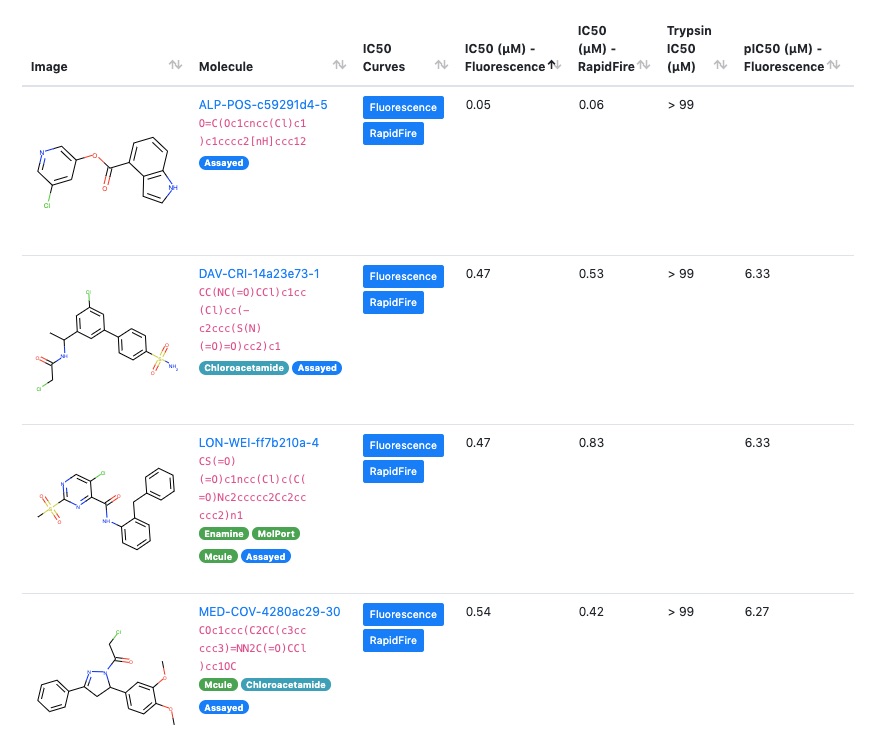
Because the amazing DiamondMX/XChem beamline is a key part of the Moonshot collaboration, we have crystal structures for nearly all of these leads, which show precisely how the molecules bind in the active site to block the viral life cycle.
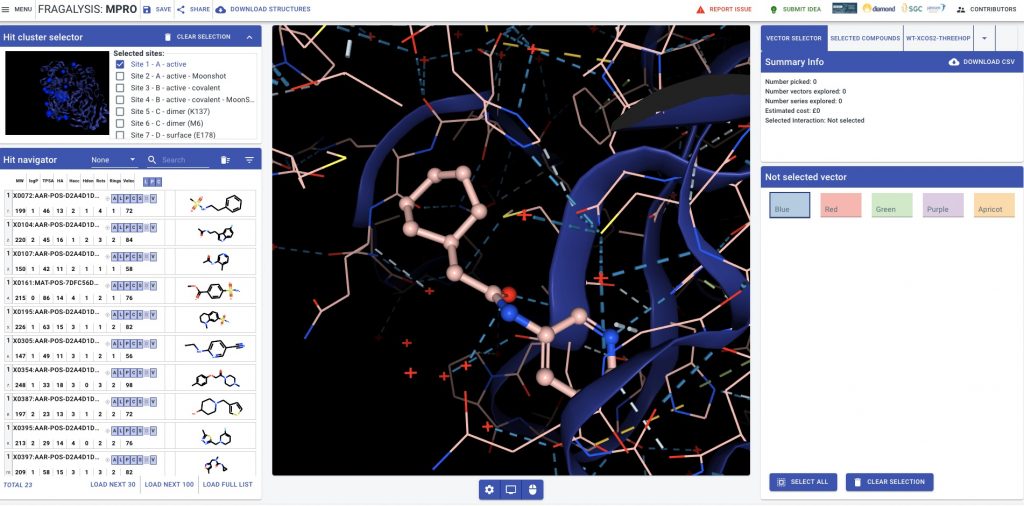
These structures are solved just a few days after the compounds are made, making it possible for us at Folding@home to use these structures to model in new compounds the chemists could make to grow the lead compounds into various pockets that could make the compounds more potent and more active in viral assays. This means we can use the awesome power of Folding@home to run thousands of alchemical free energy calculations per week to help the chemists pick which molecules to make.
While the name may sound silly, these calculations use accurate physical models (like the Open Force Field Initiative small molecule force fields) and rigorous statistical mechanics to compute whether a proposed molecule that adds a few atoms will bind better or worse than the previous design, and by how much. Our calculations use nonequilibrium trajectories to recover equilibrium free energies, using concepts pioneered by Gavin Crooks in his PhD thesis in 1999 that are now fully developed into state-of-the-art techniques for structure-based drug design.
We need your help: Weekly sprints
Here’s the part where we need your help: These calculations are enormously expensive, requiring millions of work units to run quickly on GPUs to get the chemists answer in just a couple of days. As COVID-19 makes its way around the world, it’s clear that we need to speed up the design process as much as humanly possible so we can close the cycle time between collecting new data and generating new designs that get us closer to clinical trials.
That’s why we’re introducing weekly sprints: Each week, thousands of new designs from the chemists will go up on Folding@home as part of the 134xx project series, and we’ll need your help to crunch through the hundreds of thousands of work units as quickly as possible so we can get the chemistry design team data for making the best decisions during their weekly design meetings. All the molecules that are ordered, assayed, and crystallized will be posted back to the COVID Moonshot site as soon as it is collected, which is also feeding other drug discovery projects around the world with its open science, patent-free approach to drug discovery. All our Folding@home data will also go into an open public dataset through the AWS Public Dataset Program for other computational researchers to explore.
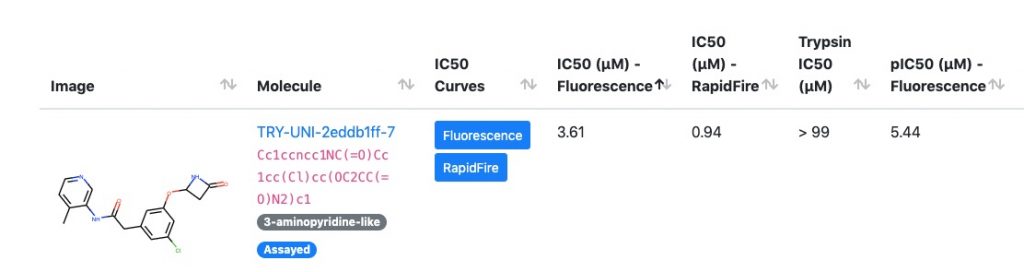
The first sprint begins right now, with thousands of primary amine elaborations and hundreds of boronic ester enumerations on our 3-aminopyridine lead that aim to fill out the S4 pocket. These are running right now, and the more GPUs we have running Folding@home, the faster we can get the chemists the answers they need. So rev up your GPU rigs and get folding!
Open source, open science
As always, all our scripts and code are open source and online, and all the data is available on the COVID Moonshot or through the MolSSI COVID-19 Structure and Therapeutics Hub we helped to build. We’re working to refine our tools to better scale up to the throughput we need to sustain and support the Moonshot as it pushes toward potency in viral assays.
Help us help you
Want to help the effort in addition to lending your GPU power? Consider donating to COVID Moonshot fundraiser to help fund the synthesis and testing of more compounds—all data is posted online as soon as it is measured, and will help us work toward a patent-free inexpensive therapy.

