In projects 10491-10499, the Chodera lab takes a look at mTOR, a serine/threonine kinase. The MTOR gene was originally discovered in yeast in 1991 and named TOR1/2 because it was the target of rapamycin, an anti-fungal small molecule isolated from the soil of Easter Island in the 1970s. In 1994, the mammalian target of rapamycin (mTOR) was discovered by Drs. Sabatini, Snyder, Abraham, and Schreiber.
 mTOR integrates multiple signal inputs to control processes such as cell growth and metabolism, among others. Due to its role in controlling a number of cellular processes, mTOR has clinical significance in neurodegenerative diseases, diabetes and cancer. In the Chodera lab, we are working with the Hsieh lab at MSKCC to understand mTOR’s role in cancer and the development of new and better therapeutics that target it.
mTOR integrates multiple signal inputs to control processes such as cell growth and metabolism, among others. Due to its role in controlling a number of cellular processes, mTOR has clinical significance in neurodegenerative diseases, diabetes and cancer. In the Chodera lab, we are working with the Hsieh lab at MSKCC to understand mTOR’s role in cancer and the development of new and better therapeutics that target it.
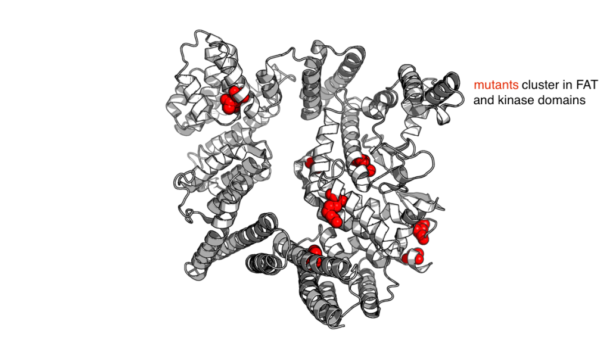
Currently, the FDA has approved treatment for metastatic clear cell Renal Cell Carcinoma (ccRCC) that includes mTOR inhibitors such as Everolimus and Temsirolimus. An effort to understand the patient-to-patient variation in response to these drugs by studying how extraordinary responders lead to the characterization of mTOR activating missense mutations in these patients. These mutants cluster in two domains of mTOR: the kinase and FAT domains. These projects will allow us to generate a model of the conformations available to mTOR and ultimately to investigate how these clinically relevant mutations might influence the protein’s structure.
Both the mTOR kinase domain and the larger construct including the FAT domain are very large systems, exploiting the latest OpenMM GPU core (0x21) and push the capabilities of latest-generation GPUs to their full extent.