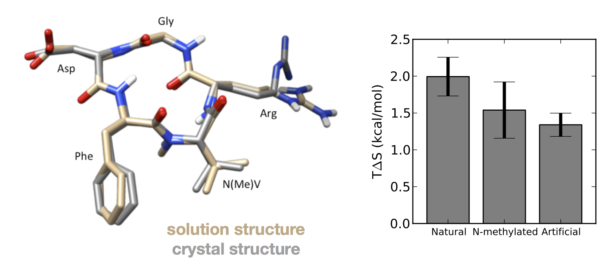
RGD peptides (those which contain a conserved Arg-Gly-Asp sequence) can adopt a hairpin loop structure found in integrin-binding proteins such as fibronectin, fibrinogen, and vitronectin. Over the last few decades, researchers have sought to develop cyclic RGD peptides that can bind potently to integrin, an important cancer target. The most successful of these to date, cilengitide (cyclo-RGDf-[N-Me]V), was discovered using a strategy of screening various chemical substitutions to find designs whose structures were more rigid. Cilengitide inhibits the binding of tumor cells to the extracellular matrix, which prevents vascularization and induces cell apoptosis. While cilengitide in particular has performed disappointingly in recent phase III clinical trials against glioblastoma, cyclic peptides in general are an exciting new platform for drug development.
In several projects soon to be released to Folding@home, we are trying to understand how to design rigid cyclic peptides in silico. First, we are have been quantifying just how accurate molecular simulations are at predicting the conformational properties of cyclic RGD peptides in solution. In a submitted manuscript, we report excellent agreement with previously published experimental data. Second, we are simulating the binding of cyclic RGD peptides to integrin to understand the relative contributions of conformational entropy (i.e. rigidity) and enthalpy (i.e. the strength of the intermolecular binding forces). Several previous studies have shown that achieving the right balance of these factors is tricky—a problem we hope our work can address.