Steven Ryckbosch, a graduate student in the Pande Group recently presented his work on Bryostatin. Folding@home projects 9000-9015 are running simulations to help answer the questions he has about it’s structure and function.
Bryostatin is a naturally occurring marine molecule that shows promising and unique activity against several diseases (most notably, cancer, HIV/AIDS, HIV latency, and Alzheimer’s). Its main target, protein kinase C (PKC), is a signaling protein central to many cellular functions. In its active form, PKC binds its ligand and is associated with the cell membrane, but we currently lack structural information about this complex in its membrane microenvironment.
The simulations performed on FAH will help to provide a structure to the PKC-ligand-membrane complex. This is complicated by the fact that while other compounds such as the phorbol esters also bind to PKC, they exhibit extremely different effects in cells and organisms. The structure and dynamics of this complex would allow us to understand bryostatin and other ligands’ binding mode and thus how to modify and tune it’s structure to improve function or even create new functions as needed for new therapies in the clinic.
Some questions Steven and the group are trying to answer:
How can we use simulation to find protein-membrane structures?
How can ligands modulate protein-membrane interaction?
How are membranes affecting bryostatin function?
How can this inform our design of new bryostatin analogues?
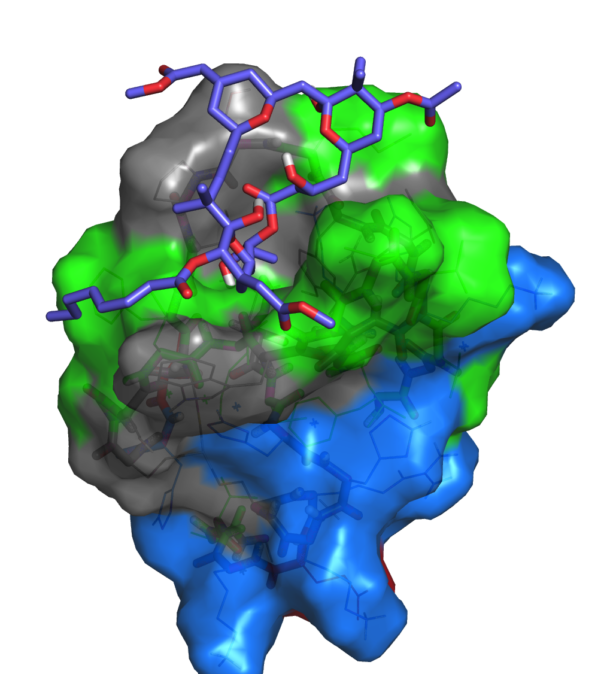
Below is a molecular simulation model of bryostatin bound to PKC’s active site.